Conclusion/Future Directions
What is Aortic Valve Disease 1?
|
AoVD1 is an autosomal dominant congenital malformation (with incomplete penetrance) of the aortic valve which connects the heart with the rest of the body. Defects in this valve prevent proper blood flow and can have serious health impacts. The character on the right in is looking directly at the aorta (in red) with his magnifying glass.
|
What is my focus?
|
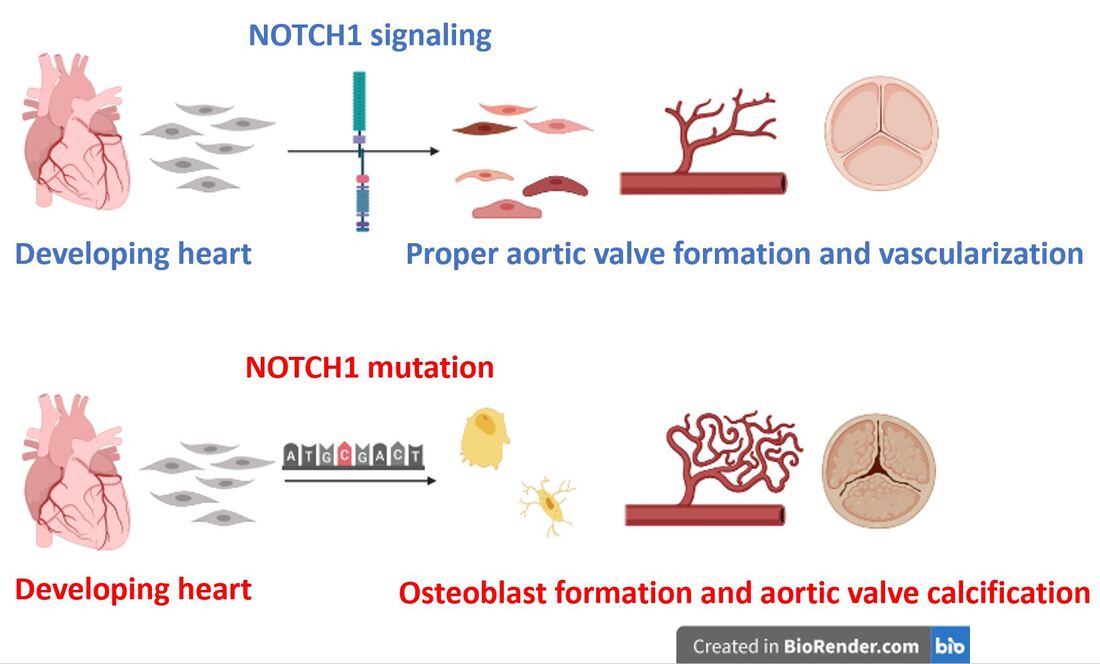
While AoVD1 manifests in several different phenotypes, I will focus on aortic valve calcification in which calcium build up on the valve leads to hardening and impaired function. This phenotype results from improper differentiation during valve development and a shift from smooth muscle production to osteoblast (the major cell type found in bones) production.
|
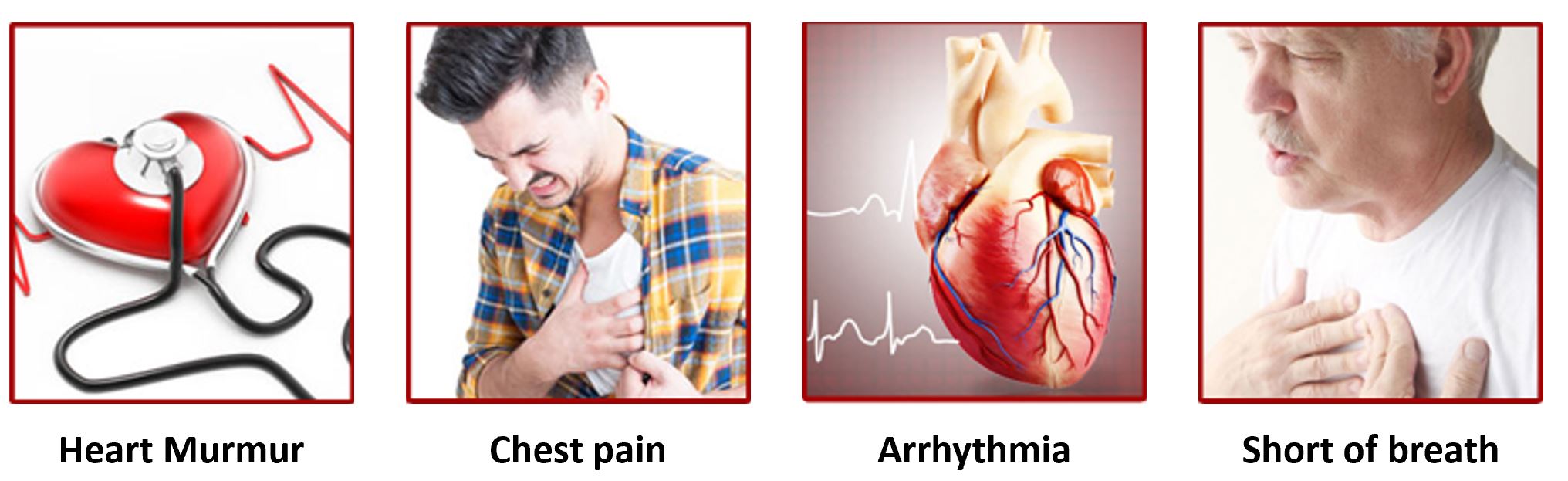
What are the symptoms of AoVD1?
Why did I pick AoVD1?
What is NOTCH1?
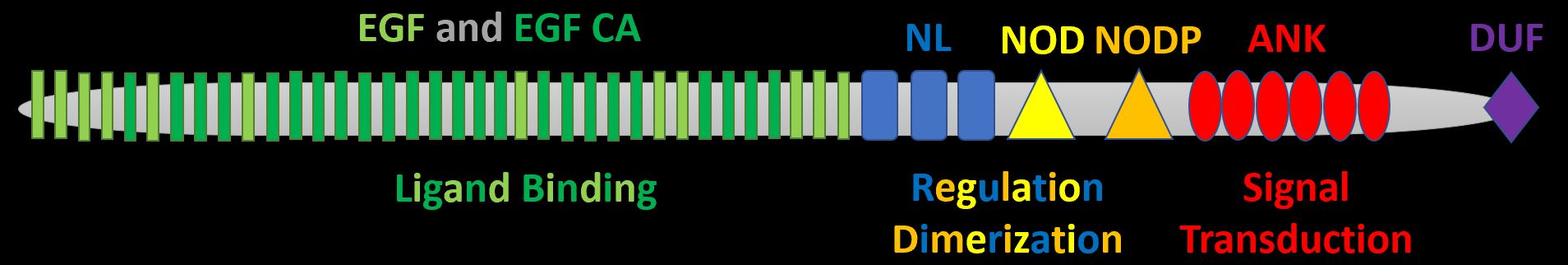
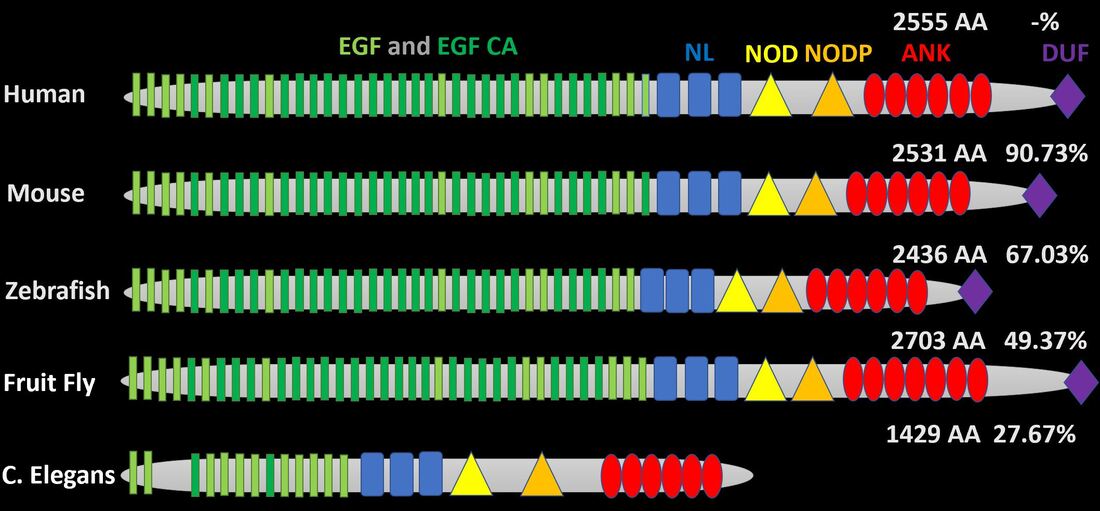
The most notable association with AoVD1 is the transmembrane receptor NOTCH1. It possesses N-terminal EGF-like and EGF-like calcium binding domains responsible for ligand binding, NL, NOD, and NODP domains responsible for protein regulation and dimerization, and ANK domains responsible for signal transduction. There is also a C-terminal domain of unknown function.
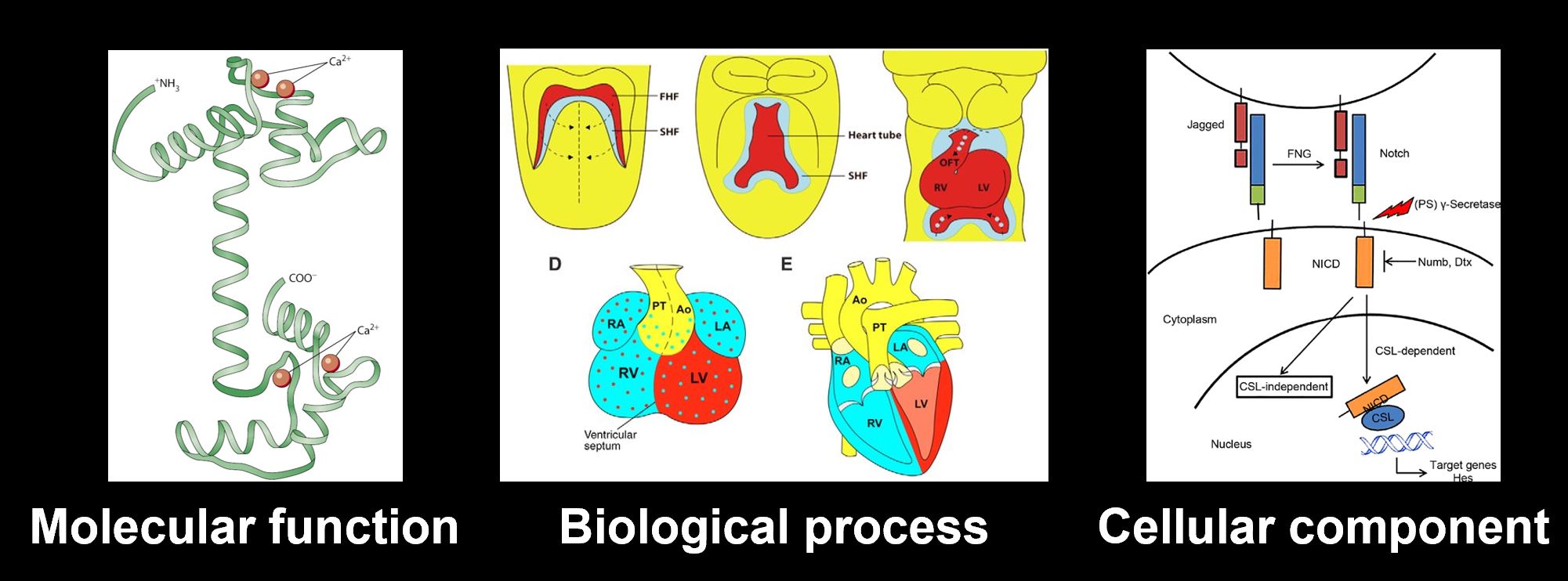
What gene ontologies will guide my research?
What is NOTCH signaling?
|
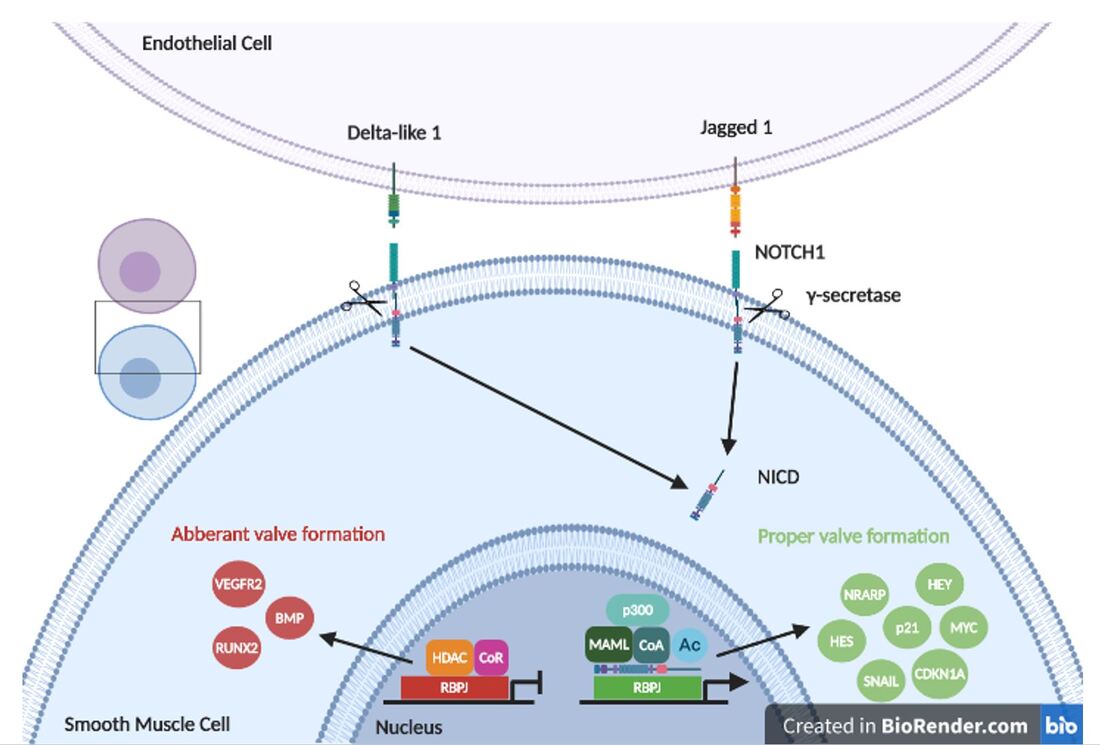
NOTCH signaling is key to proper cell fate determination in the heart. NOTCH1 is stimulated by Jag1 or Delta-like 1 and then cleaved by gamma secretase. Its intracellular domain then translocates to the nucleus to interact with epigenetic factors and transcirptional co-regulators to stimulate transcriptional activation of smooth muscle enhancers like HEY and HES. Loss of NOTCH1 signaling leads to upregulation of osteoblast factors like RUNX2.
|
How well is NOTCH1 conserved?
|
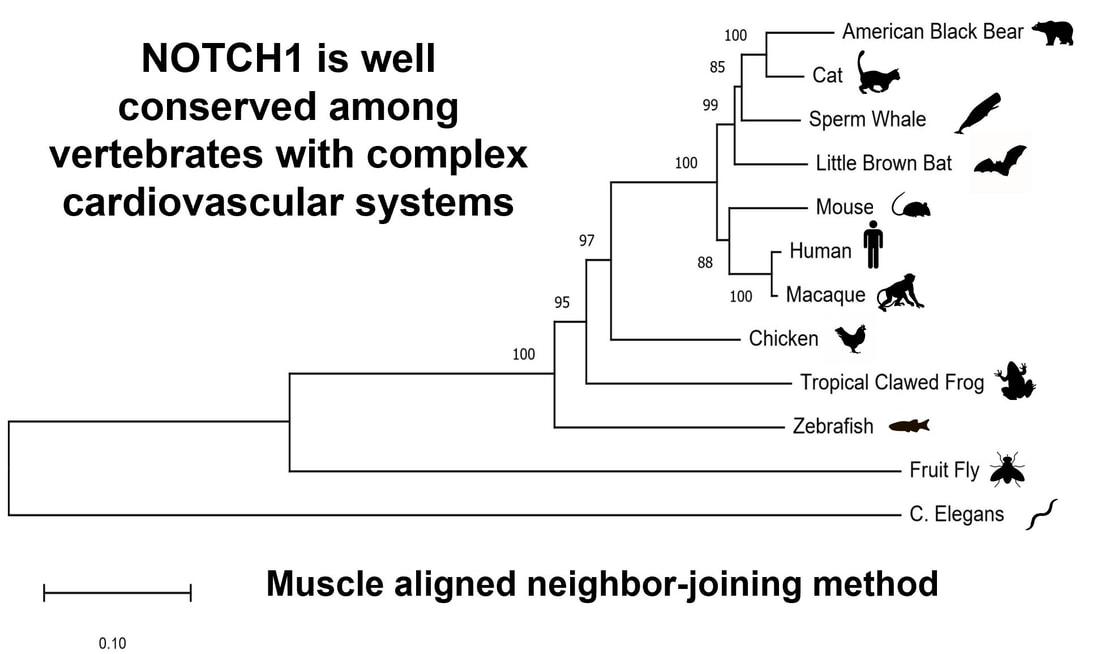
NOTCH1 is well conserved in organisms with complex cardiovascular systems (hearts). The EGF-like and EGF-like calcium binding domains are especially conserved in animals with hearts as humans have 36, mice have 36, zebrafish have 35, Drosophila have 37, and C. elegans, which has a more simple, open cardiovascular system, only has 13. Through phylogenetic analysis we can see that C. elegans is the clear out-group. |
Can zebrafish be used to study AoVD1?
|
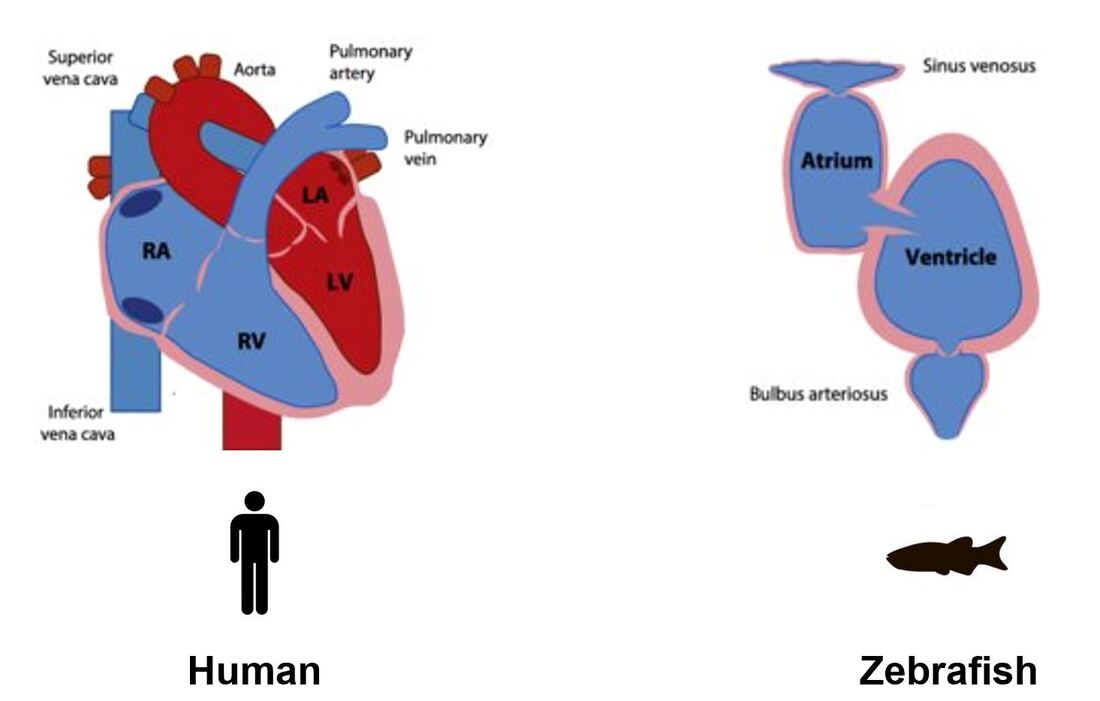
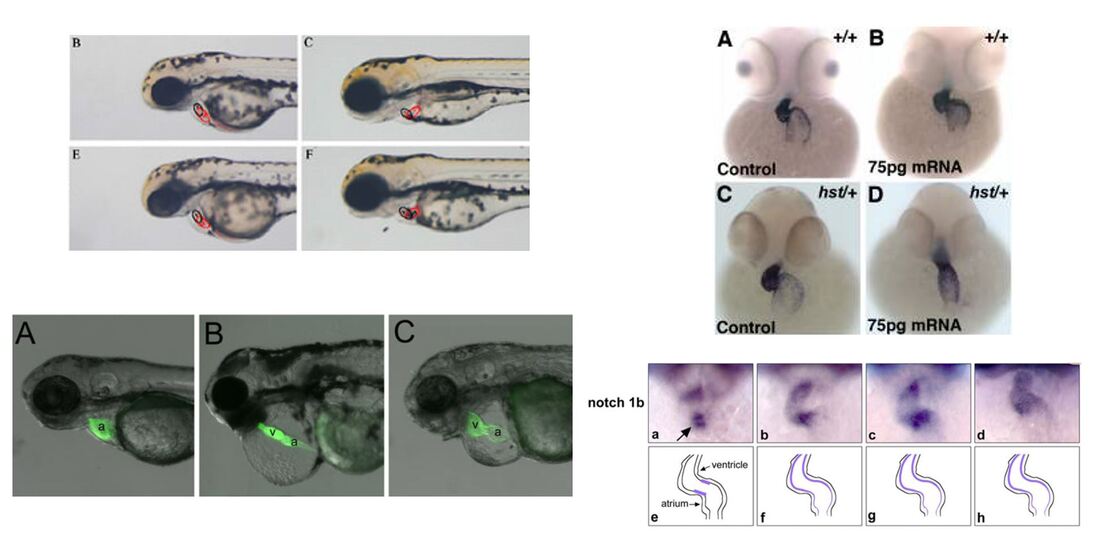
Although zebrafish do not have four chambered hearts like humans, other mammals, and birds, their two chambered architecture allows for visualization of valve formation between their atrium and ventricle. In addition to similar cardiovascular systems, zebrafish have easily manipulated genomes and aortic valve formation can be monitored by staining, GFP, or simply by naked eye. |
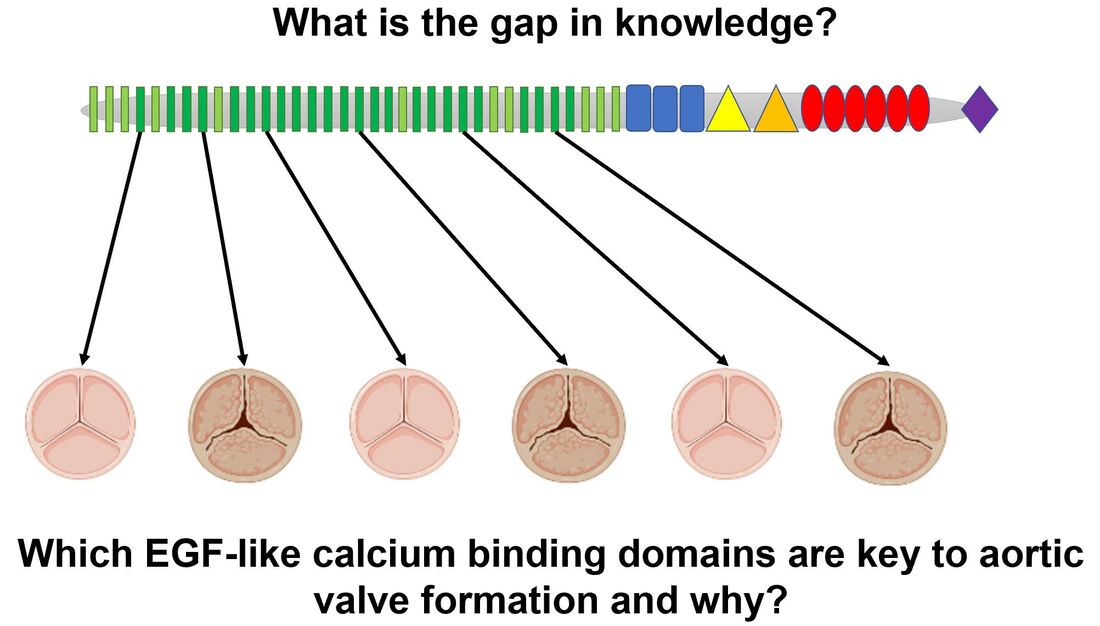
What is the gap in knowledge?
|
In 2004, Cripe et al. showed that AoVD1 is heritable [1]. In 2005, Garg et al. showed that NOTCH1 was responsible for this inheritance [2]. Since then the molecular framework I have presented above has been pieced together; however, it remains to be seen which EGF-like calcium binding domains are necessary for proper aortic valve formation and why.
|
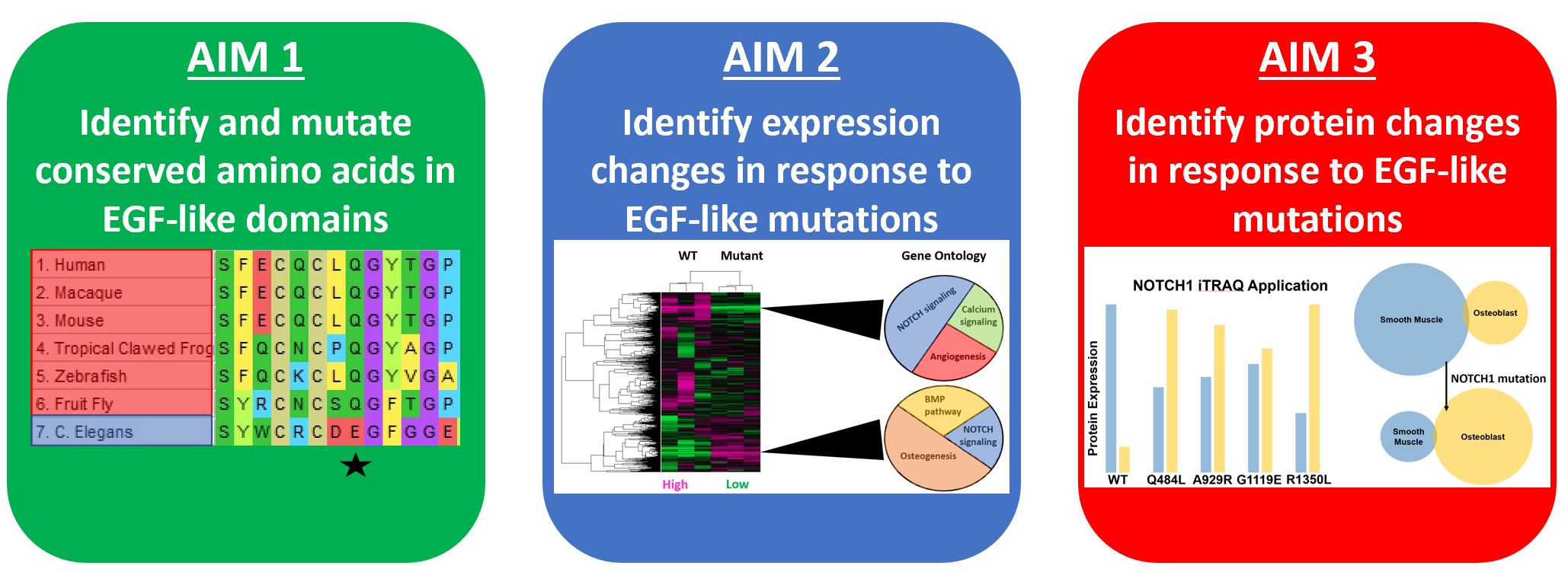
What are my aims?
|
To accomplish this I will determine which EGF-like calcium binding domains are conserved in organisms with hearts, mutate amino acids conserved in these domains, and attempt to observe the shift from smooth muscle differentiation to obsteoblast differentiation in both gene expression and protein expression.
|
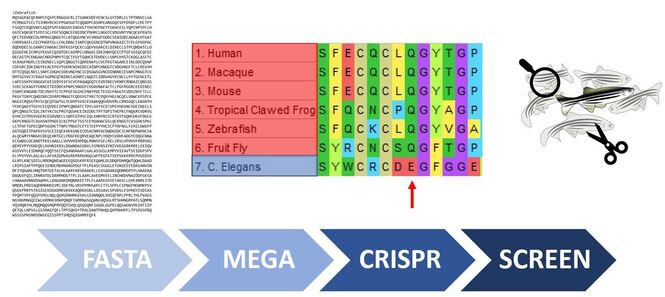
Hypothesis 1
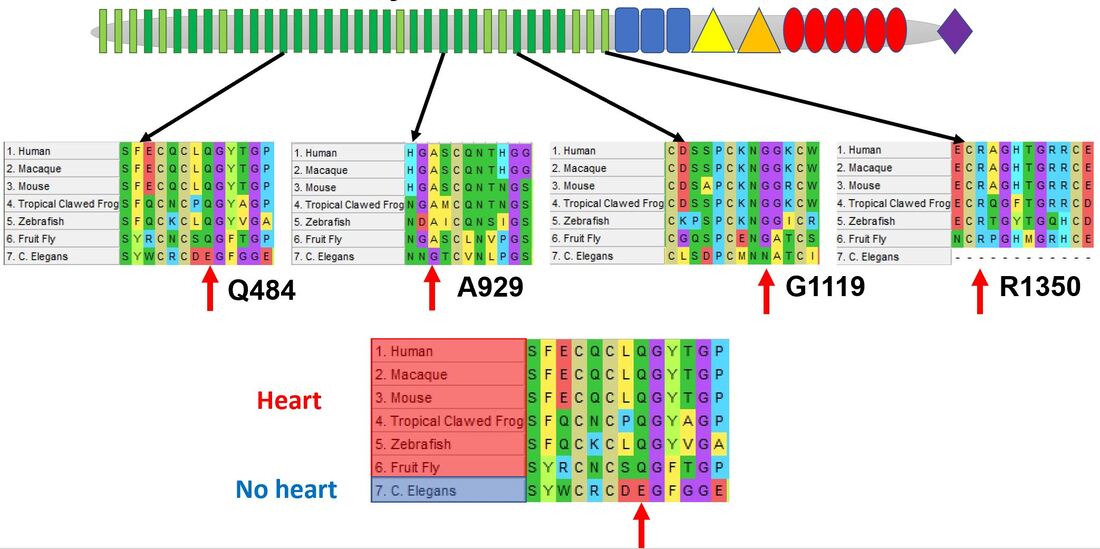
Mutation of EGF-like calcium binding domains conserved between model organism with a heart will lead to aortic valve calcification: I will obtain NOTCH1 FASTA sequences of common model organisms, align in MEGA, and identify amino acids within EGF-like calcium binding domains conserved in animals with hearts but not in more simple organisms. I will then mutate with CRISPR-cas9 and screen for aortic valve calcification with the naked eye, confirming calcification by staining for osteoblast markers.
Aim 1
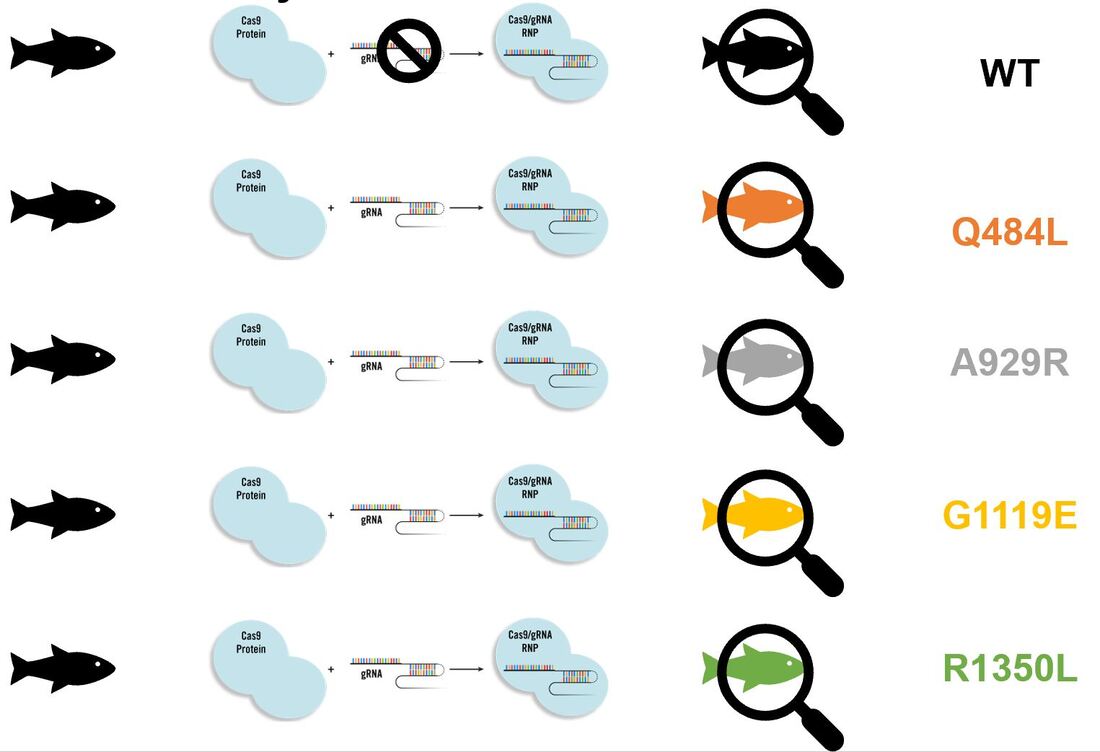
Determine which EGF-like calcium binding domain in NOTCH1 is necessary for aortic valve calcification: The first three alignments display amino acids conserved in organisms with hearts while the fourth is a common mutation found in clinic. Because NOTCH1 is a transmembrane protein I will test out a host of mutation types (for example Q484L is a polar to non-polar shift while A929R is a non-polar to positively charged shift).
Hypothesis 2
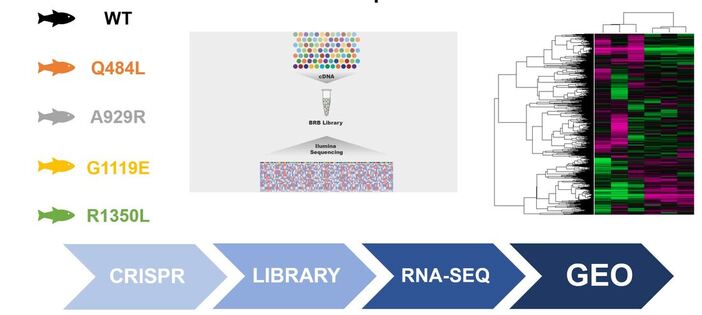
Mutation of conserved EGF-like calcium binding regions will lead to a shift from gene expression highly enriched in smooth muscle formation to gene expression highly enriched in osteogenesis: I will isolate aortic valves from the lines created in Aim 1, extract RNA, perform RNA-seq, and publish my results on GEO for other researchers to reference.
Aim 2
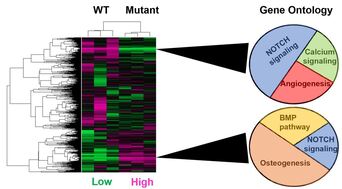
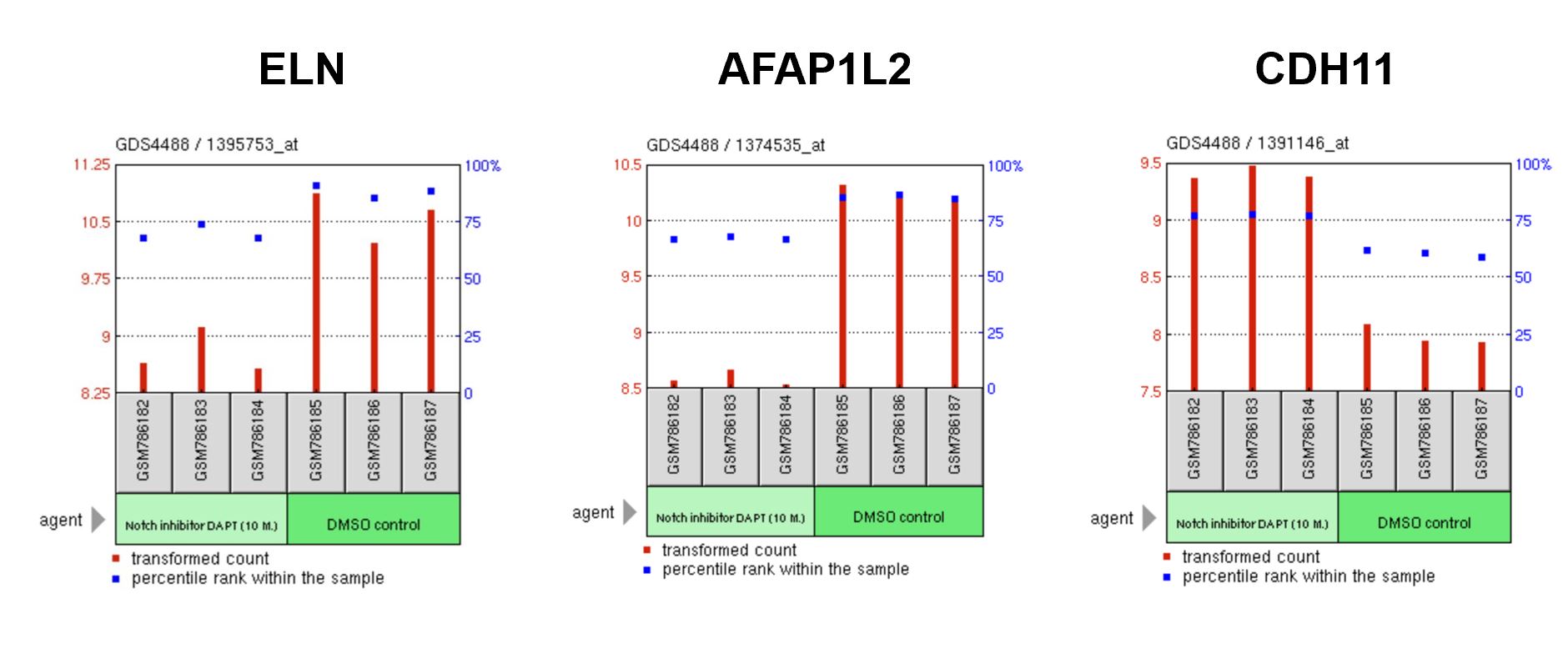
Identify which EGF-like calcium binding domains are important for maintaining gene expression related to smooth muscle differentiation: The figures below are gene expression arrays from the Acharya et al. study which used siRNA to inhibit NOTCH1 signaling. I would expect to see an overall shift from NOTCH and calcium signaling as well as angiogenesis to ontologies like osteogenesis and the bone morphogenic protein pathway. In addition, individual heart-specific genes like elastin (ELN) and actin filament associated protein (AFAP1L2) will be downregulated while osteoblast-specific genes like cadherins (CDH11) will be upregulated.
Hypothesis 3
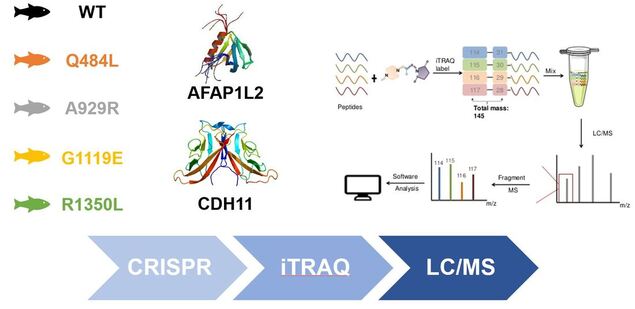
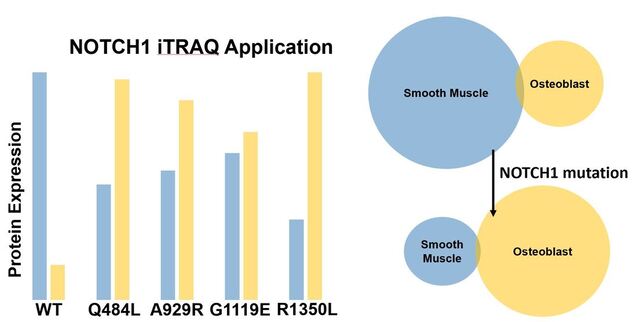
Mutation of conserved EGF-like calcium binding regions will lead to a shift from protein expression highly enriched in smooth muscle formation to protein expression highly enriched in osteogenesis: Using my lines created in Aim 1 I will isolate aortic valves, label each line with isobaric tags specific to a smooth muscle-specific marker like AFAP1L2 and an osteoblast-specific marker like CDH11, and then analyze by way of liquid chromatography/mass spectrometry.
Aim 3
Identify which EGF-like calcium binding domains are important for maintaining protein expression responsible for smooth muscle differentiation: I expect to see a protein expression pattern mirroring that of the gene expression pattern in which a shift from smooth muscle expression to osteoblast expression occurs upon NOTCH1 mutation.
Future Goals
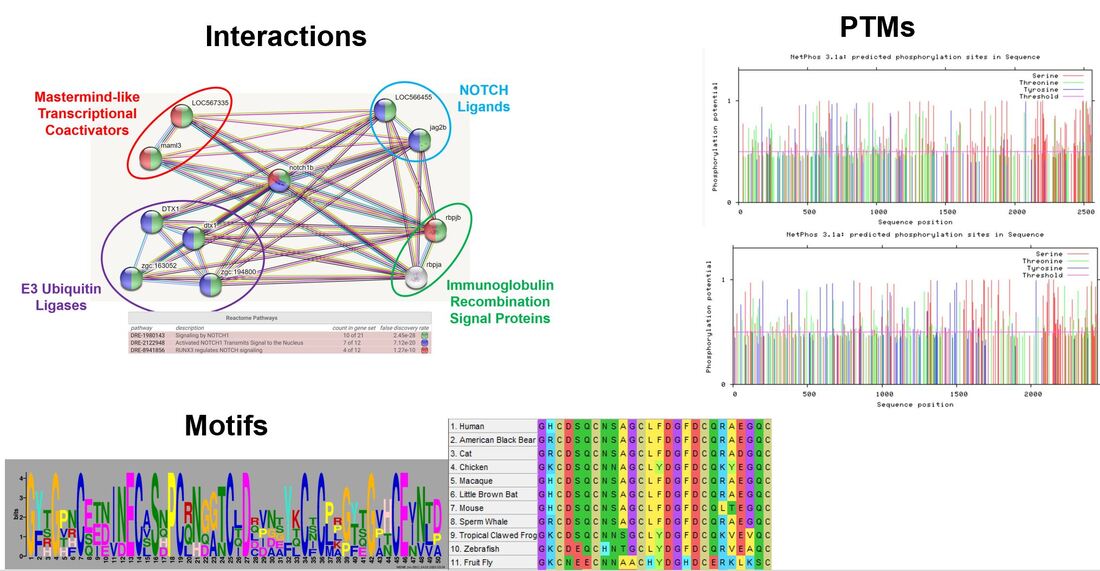
I would like to continue interrogating the EGF-like calcium binding domains starting with their role in ligand binding. I am also interested in post-translational modifications as phosphorylation and several carbohydrate modifications have been shown to play important roles in ligand preference and proper NOTCH cleavage. Finally, the EGF-like calcium binding domains contain many glycine-cysteine motifs which provide insight into its secondary structure. Disrupting these features may help us to further understand the molecular basis of AoVD1.
Final Presentation Drafts
| mario_bertogliat_final_pres_4.27.20.pdf | |
| File Size: | 2633 kb |
| File Type: | |
| mario_bertogliat_final_pres_4.27.20.pptx | |
| File Size: | 7377 kb |
| File Type: | pptx |
| mario_bertogliat_aovd1_final_pres_4.2.20.pptx | |
| File Size: | 5061 kb |
| File Type: | pptx |
| mario_bertogliat_aovd1_final_pres_2.27.20.pptx | |
| File Size: | 3374 kb |
| File Type: | pptx |
"This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison."
Last Updated: 5/3/20
www.genetics564.weebly.com
Last Updated: 5/3/20
www.genetics564.weebly.com
Images:
1.https://healthblog.uofmhealth.org/Mens-Heart-Disease-Risks-Signs-perfcon
2.https://biorender.com/
3.https://images.medindia.net/patientinfo/950_400/symptoms-signs-of-bicuspid-aortic-valve.jpg
4.https://www.balladhealth.org/transcatheter-aortic-valve-replacement-tavr
5. https://www.google.com/search?q=jesse+bertogliat&rlz=1C1EJFC_enUS811US811&source=lnms&tbm=isch&sa=X&ved=2ahUKEwj09c-5j__oAhVTHc0KHQTPB8gQ_AUoAXoECAsQAw&biw=1368&bih=753&dpr=2#imgrc=d1hVSokdboLEjM
6.https://www.google.com/search?q=julian+bertogliat&rlz=1C1EJFC_enUS811US811&source=lnms&tbm=isch&sa=X&ved=2ahUKEwjOtYDnj__oAhVTQ80KHaSzA3EQ_AUoAnoECAsQBA&biw=1368&bih=753#imgrc=s-xac8mMZX5EbM
7.http://smart.embl-heidelberg.de/
8.http://www.mun.ca/biology/desmid/brian/BIOL2060/BIOL2060-14/CB14old.html
9.Cohen ED(1), Tian Y, Morrisey EE. Wnt signaling: an essential regulator of cardiovascular differentiation, morphogenesis and progenitor self-renewal. Development. 2008 Mar;135(5):789-98. doi: 10.1242/dev.016865.
10.Hao X, Tian J, Yin L, et al. (2017). MRI detects protective effects of DAPT treatment with modulation of microglia/macrophages at subacute and chronic stages following cerebral ischemia. Molecular Medicine Reports. 16. 10.3892/mmr.2017.7200.
11. https://biorender.com/
12. http://smart.embl-heidelberg.de/
13.https://image.freepik.com/freie-ikonen/bar-seitenansicht-silhouette_318-42619.jpg
14.https://image.flaticon.com/icons/svg/47/47197.svg
15.http://imageog.flaticon.com/icons/png/512/47/47096.png?size=1200x630f&pad=10,10,10,10&ext=png&bg=FFFFFFFF
16. https://photos.gograph.com/thumbs/CSP/CSP418/bat-silhouette-stock-illustration_k19082699.jpg
17.http://imageog.flaticon.com/icons/png/512/47/47240.png?size=1200x630f&pad=10,10,10,10&ext=png&bg=FFFFFFFF
18.https://upload.wikimedia.org/wikipedia/commons/thumb/d/d8/Person_icon_BLACK-01.svg/721px-Person_icon_BLACK-01.svg.png
19.https://image.freepik.com/free-icon/monkey_318-62892.png
20.https://image.freepik.com/free-icon/chicken-symbol_318-10389.jpg
21.https://image.shutterstock.com/image-photo/image-450w-59417659.jpg
22.https://media.istockphoto.com/vectors/vector-set-of-black-aquarium-fish-silhouettes-with-text-vector-id1145856310?k=6&m=1145856310&s=612x612&w=0&h=gOnb4kwuztGEmfrxQpe4RoNNW3OkwoTyLOnIHelnWYE=
23.https://d30y9cdsu7xlg0.cloudfront.net/png/40840-200.png
24.https://imageog.flaticon.com/icons/png/512/47/47217.png?size=1200x630f&pad=10,10,10,10&ext=png&bg=FFFFFFFF
25.van Opbergen C, van der Voorn S, et al. Cardiac Ca2+ signalling in zebrafish: Translation of findings to man. Prog Biophys Mol Biol. 2018 Oct;138:45-58. doi: 10.1016/j.pbiomolbio.2018.05.002
26.https://upload.wikimedia.org/wikipedia/commons/thumb/d/d8/Person_icon_BLACK-01.svg/721px-Person_icon_BLACK-01.svg.png
27.https://media.istockphoto.com/vectors/vector-set-of-black-aquarium-fish-silhouettes-with-text-vector-id1145856310?
28.Li M, Hu X, et al. Overexpression of miR-19b Impairs Cardiac Development in Zebrafish by Targeting ctnnb1. Cell Physiol Biochem 2014;33:1988-2002; https://doi.org/10.1159/000362975
29.Lee YM, Cope JJ, Ackermann GE, et al. Vascular endothelial growth factor receptor signaling is required for cardiac valve formation in zebrafish. Dev Dyn. 2006;235(1):29–37. doi:10.1002/dvdy.20559
30.Ribeiro I, Kawakami Y, Büscher D, et al. Tbx2 and Tbx3 regulate the dynamics of cell proliferation during heart remodeling. PLoS One. 2007;2(4):e398. Published 2007 Apr 25. doi:10.1371/journal.pone.0000398
31.Camarata T, Krcmery J, et al. Pdlim7 (LMP4) regulation of Tbx5 specifies zebrafish heart atrio-ventricular boundary and valve formation. Dev Biol. 2010;337(2):233–245. doi:10.1016/j.ydbio.2009.10.039
32.http://smart.embl-heidelberg.de/
33.https://biorender.com/
34.Acharya A, Hans CP, Koenig SN, et al. Inhibitory role of Notch1 in calcific aortic valve disease. PLoS One. 2011;6(11):e27743. doi:10.1371/journal.pone.0027743
35.https://www.genome.gov/sites/default/files/tg/en/illustration/zebrafish.jpg
36.http://smart.embl-heidelberg.de/
37.https://www.mirusbio.com/assets/figures/crispr-rnp-delivery-approach.png
38.https://phys.org/news/2019-04-brb-seq-quick-cheaper-future-rna.html
39.Acharya A, Hans CP, Koenig SN, et al. Inhibitory role of Notch1 in calcific aortic valve disease. PLoS One. 2011;6(11):e27743. doi:10.1371/journal.pone.0027743
40.Acharya A, Hans CP, Koenig SN, et al. Inhibitory role of Notch1 in calcific aortic valve disease. PLoS One. 2011;6(11):e27743. doi:10.1371/journal.pone.0027743
41. Acharya A, Hans CP, Koenig SN, et al. Inhibitory role of Notch1 in calcific aortic valve disease. PLoS One. 2011;6(11):e27743. doi:10.1371/journal.pone.0027743
42.https://www.phosphosite.org/proteinAction?id=4755&showAllSites=true
43.https://en.wikipedia.org/wiki/CDH11
44.https://www.slideshare.net/Creative-Proteomics/mass-spectrometrybased-proteomics-quantification-itraq
45.https://string-db.org/cgi/network.pl?taskId=GEuQXPpvYjra
46. http://www.cbs.dtu.dk/services/NetPhos/
47.http://meme-suite.org/
Information:
1.Linda Cripe, Gregor Andelfinger, Lisa J. Martin, Kerry Shooner, D.Woodrow Benson, Bicuspid aortic valve is heritable, Journal of the American College of Cardiology, Volume 44, Issue 1, 2004, Pages 138-143, ISSN 0735-1097.
2. Garg, V., Muth, A., Ransom, J. et al. Mutations in NOTCH1 cause aortic valve disease. Nature 437, 270–274 (2005).
3.Acharya A, Hans CP, Koenig SN, et al. Inhibitory role of Notch1 in calcific aortic valve disease. PLoS One. 2011;6(11):e27743. doi:10.1371/journal.pone.0027743
1.https://healthblog.uofmhealth.org/Mens-Heart-Disease-Risks-Signs-perfcon
2.https://biorender.com/
3.https://images.medindia.net/patientinfo/950_400/symptoms-signs-of-bicuspid-aortic-valve.jpg
4.https://www.balladhealth.org/transcatheter-aortic-valve-replacement-tavr
5. https://www.google.com/search?q=jesse+bertogliat&rlz=1C1EJFC_enUS811US811&source=lnms&tbm=isch&sa=X&ved=2ahUKEwj09c-5j__oAhVTHc0KHQTPB8gQ_AUoAXoECAsQAw&biw=1368&bih=753&dpr=2#imgrc=d1hVSokdboLEjM
6.https://www.google.com/search?q=julian+bertogliat&rlz=1C1EJFC_enUS811US811&source=lnms&tbm=isch&sa=X&ved=2ahUKEwjOtYDnj__oAhVTQ80KHaSzA3EQ_AUoAnoECAsQBA&biw=1368&bih=753#imgrc=s-xac8mMZX5EbM
7.http://smart.embl-heidelberg.de/
8.http://www.mun.ca/biology/desmid/brian/BIOL2060/BIOL2060-14/CB14old.html
9.Cohen ED(1), Tian Y, Morrisey EE. Wnt signaling: an essential regulator of cardiovascular differentiation, morphogenesis and progenitor self-renewal. Development. 2008 Mar;135(5):789-98. doi: 10.1242/dev.016865.
10.Hao X, Tian J, Yin L, et al. (2017). MRI detects protective effects of DAPT treatment with modulation of microglia/macrophages at subacute and chronic stages following cerebral ischemia. Molecular Medicine Reports. 16. 10.3892/mmr.2017.7200.
11. https://biorender.com/
12. http://smart.embl-heidelberg.de/
13.https://image.freepik.com/freie-ikonen/bar-seitenansicht-silhouette_318-42619.jpg
14.https://image.flaticon.com/icons/svg/47/47197.svg
15.http://imageog.flaticon.com/icons/png/512/47/47096.png?size=1200x630f&pad=10,10,10,10&ext=png&bg=FFFFFFFF
16. https://photos.gograph.com/thumbs/CSP/CSP418/bat-silhouette-stock-illustration_k19082699.jpg
17.http://imageog.flaticon.com/icons/png/512/47/47240.png?size=1200x630f&pad=10,10,10,10&ext=png&bg=FFFFFFFF
18.https://upload.wikimedia.org/wikipedia/commons/thumb/d/d8/Person_icon_BLACK-01.svg/721px-Person_icon_BLACK-01.svg.png
19.https://image.freepik.com/free-icon/monkey_318-62892.png
20.https://image.freepik.com/free-icon/chicken-symbol_318-10389.jpg
21.https://image.shutterstock.com/image-photo/image-450w-59417659.jpg
22.https://media.istockphoto.com/vectors/vector-set-of-black-aquarium-fish-silhouettes-with-text-vector-id1145856310?k=6&m=1145856310&s=612x612&w=0&h=gOnb4kwuztGEmfrxQpe4RoNNW3OkwoTyLOnIHelnWYE=
23.https://d30y9cdsu7xlg0.cloudfront.net/png/40840-200.png
24.https://imageog.flaticon.com/icons/png/512/47/47217.png?size=1200x630f&pad=10,10,10,10&ext=png&bg=FFFFFFFF
25.van Opbergen C, van der Voorn S, et al. Cardiac Ca2+ signalling in zebrafish: Translation of findings to man. Prog Biophys Mol Biol. 2018 Oct;138:45-58. doi: 10.1016/j.pbiomolbio.2018.05.002
26.https://upload.wikimedia.org/wikipedia/commons/thumb/d/d8/Person_icon_BLACK-01.svg/721px-Person_icon_BLACK-01.svg.png
27.https://media.istockphoto.com/vectors/vector-set-of-black-aquarium-fish-silhouettes-with-text-vector-id1145856310?
28.Li M, Hu X, et al. Overexpression of miR-19b Impairs Cardiac Development in Zebrafish by Targeting ctnnb1. Cell Physiol Biochem 2014;33:1988-2002; https://doi.org/10.1159/000362975
29.Lee YM, Cope JJ, Ackermann GE, et al. Vascular endothelial growth factor receptor signaling is required for cardiac valve formation in zebrafish. Dev Dyn. 2006;235(1):29–37. doi:10.1002/dvdy.20559
30.Ribeiro I, Kawakami Y, Büscher D, et al. Tbx2 and Tbx3 regulate the dynamics of cell proliferation during heart remodeling. PLoS One. 2007;2(4):e398. Published 2007 Apr 25. doi:10.1371/journal.pone.0000398
31.Camarata T, Krcmery J, et al. Pdlim7 (LMP4) regulation of Tbx5 specifies zebrafish heart atrio-ventricular boundary and valve formation. Dev Biol. 2010;337(2):233–245. doi:10.1016/j.ydbio.2009.10.039
32.http://smart.embl-heidelberg.de/
33.https://biorender.com/
34.Acharya A, Hans CP, Koenig SN, et al. Inhibitory role of Notch1 in calcific aortic valve disease. PLoS One. 2011;6(11):e27743. doi:10.1371/journal.pone.0027743
35.https://www.genome.gov/sites/default/files/tg/en/illustration/zebrafish.jpg
36.http://smart.embl-heidelberg.de/
37.https://www.mirusbio.com/assets/figures/crispr-rnp-delivery-approach.png
38.https://phys.org/news/2019-04-brb-seq-quick-cheaper-future-rna.html
39.Acharya A, Hans CP, Koenig SN, et al. Inhibitory role of Notch1 in calcific aortic valve disease. PLoS One. 2011;6(11):e27743. doi:10.1371/journal.pone.0027743
40.Acharya A, Hans CP, Koenig SN, et al. Inhibitory role of Notch1 in calcific aortic valve disease. PLoS One. 2011;6(11):e27743. doi:10.1371/journal.pone.0027743
41. Acharya A, Hans CP, Koenig SN, et al. Inhibitory role of Notch1 in calcific aortic valve disease. PLoS One. 2011;6(11):e27743. doi:10.1371/journal.pone.0027743
42.https://www.phosphosite.org/proteinAction?id=4755&showAllSites=true
43.https://en.wikipedia.org/wiki/CDH11
44.https://www.slideshare.net/Creative-Proteomics/mass-spectrometrybased-proteomics-quantification-itraq
45.https://string-db.org/cgi/network.pl?taskId=GEuQXPpvYjra
46. http://www.cbs.dtu.dk/services/NetPhos/
47.http://meme-suite.org/
Information:
1.Linda Cripe, Gregor Andelfinger, Lisa J. Martin, Kerry Shooner, D.Woodrow Benson, Bicuspid aortic valve is heritable, Journal of the American College of Cardiology, Volume 44, Issue 1, 2004, Pages 138-143, ISSN 0735-1097.
2. Garg, V., Muth, A., Ransom, J. et al. Mutations in NOTCH1 cause aortic valve disease. Nature 437, 270–274 (2005).
3.Acharya A, Hans CP, Koenig SN, et al. Inhibitory role of Notch1 in calcific aortic valve disease. PLoS One. 2011;6(11):e27743. doi:10.1371/journal.pone.0027743