NOTCH1 Gene
What is transcriptomics?
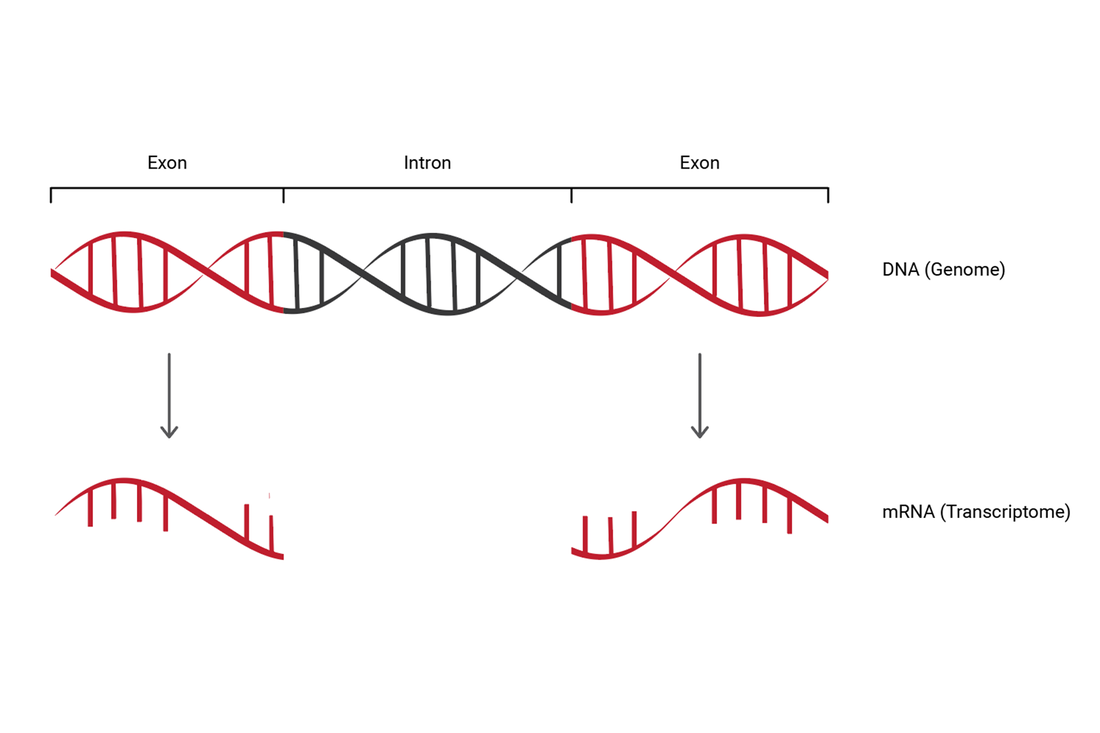
Transcriptomics is the study of the transcriptome which reflects all RNA expressed in a cell [1]. Although mRNA is the object of most transcriptomic studies, changes in other RNA like long noncoding RNA, miRNA, rRNA, and tRNA constitute much of the transcriptome [2]. To be studied, RNA is reverse transcribed into cDNA and then analyzed by microarray, qRT-PCR, or next-generation sequencing platforms [2]. The study of transcriptomics is important in identifying gene expression changes brought about by disease, treatment, or environment, and diet. Transcriptomics is also essential in determining differential gene expression between tissues or even more specifically, between individual cells.
Where is NOTCH1 expressed?
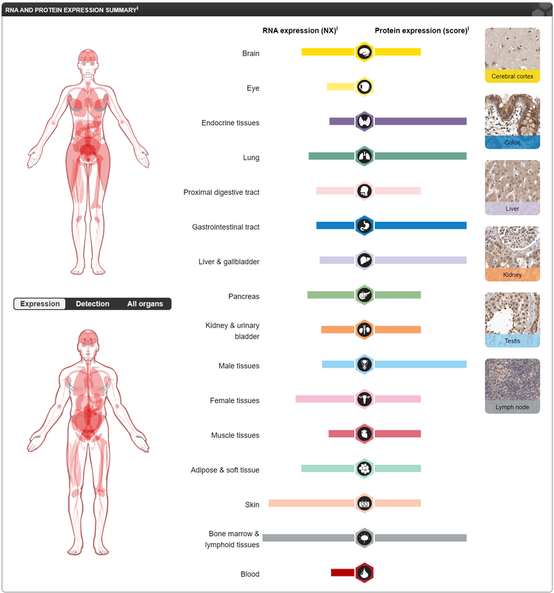
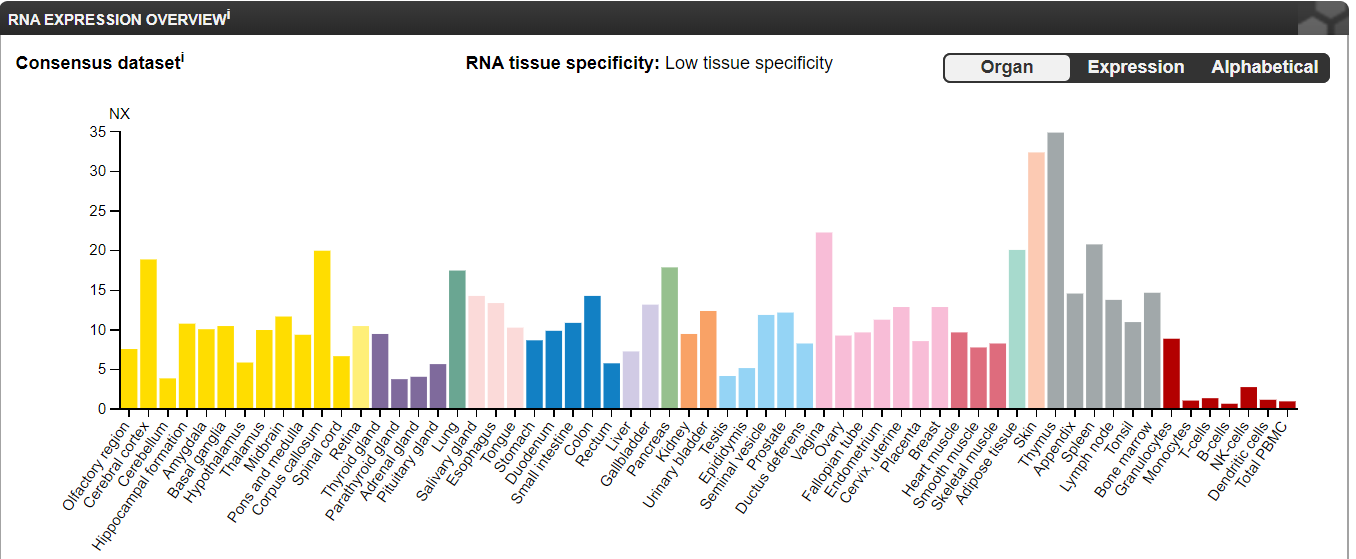
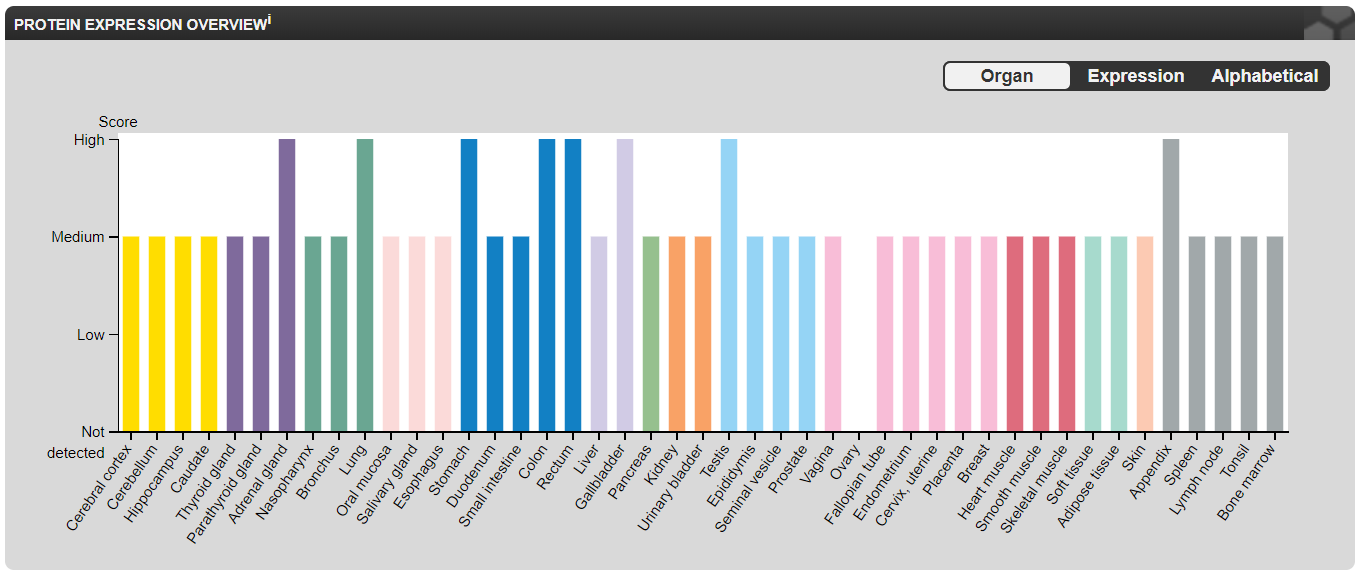
NOTCH1 lacks both RNA and protein-tissue specificity. NOTCH1 RNA expression is highest in the brain, skin, and bone marrow/lymphoid tissues while NOTCH1 protein is highly expressed in the endocrine tissues, lung, gastrointestinal tract, liver/gallbladder, male tissues, and bone marrow/lymphoid tissues [3]. Both NOTCH1 RNA and protein are significantly expressed in the heart [3]. Interestingly, NOTCH1 RNA expression is higher in female tissues but NOTCH1 protein expression is higher in male tissues [3].
NOTCH1 RNA and protein expression lacks distinct tissue-specificity [3].
Conclusion
Through transcriptomics it was found that NOTCH1 RNA and protein are expressed in a non-specific manner regarding tissue type. NOTCH1 is significantly expressed in the heart and may have sex-specific bias within transcriptional and translational programs. Because of this non-specific expression pattern, care must be taken to analyze only aortic valve tissue when studying Aortic Valve Disease 1.