NOTCH1 Protein
What are protein domains?
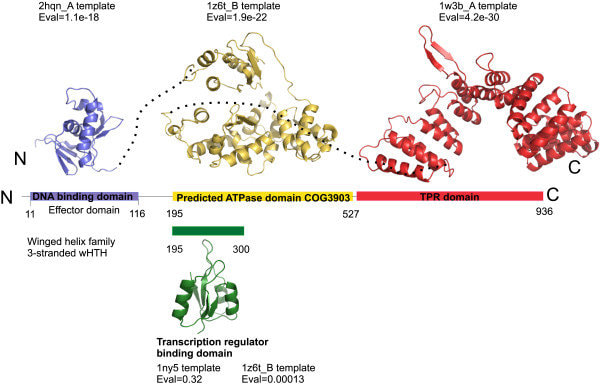
Protein domains are sequence-specific areas of homology that can be thought of as the "evolutionary unit" of protein function [1]. The have also been defined as semi-independent globular folding units in regards to protein structure [1]. A single gene may have multiple different domains that each carry a unique function. Additionally, between isoforms of the same gene, domains may arise or disappear. Motifs are similar to domains in that they represent a portion of 3D structure; however, they are found across proteins of different types and do not necessarily function on their own like domains do [2].
What domains does NOTCH1 consist of?
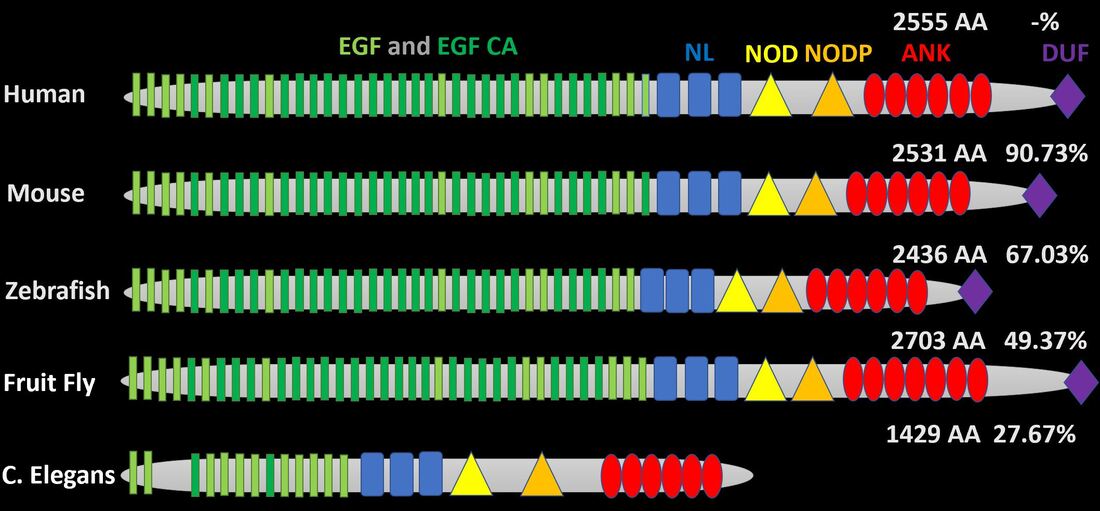
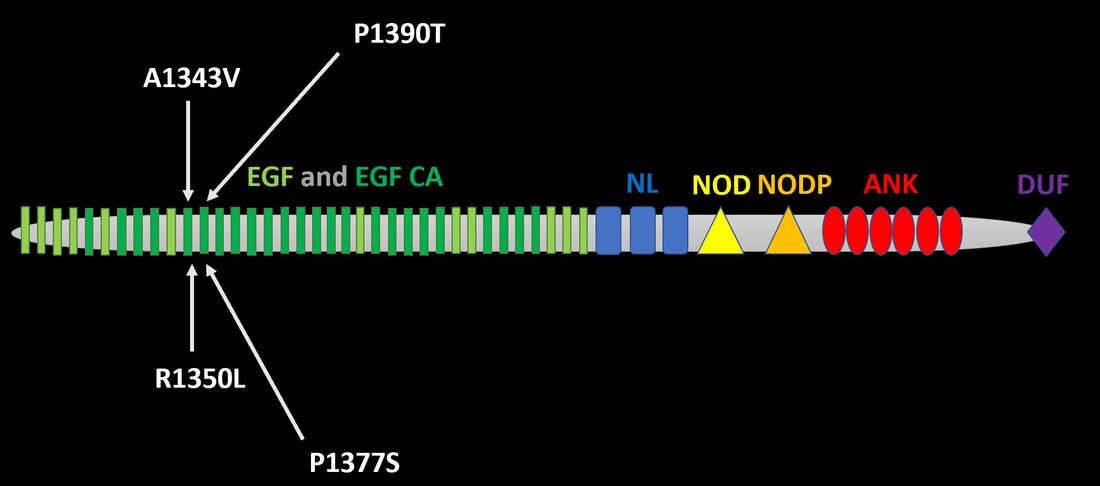
Human NOTCH1 has 36 epidermal growth factor (EGF)-like repeats with alternating calcium-binding EGF-like repeats [3]. This area of the gene is highly conserved a mice have 36, zebrafish have 35, fruit flies have 37, and C. elegans have 13 [3]. In addition, the NL, NOD, and NODP domains, which function in regulation and dimerization, are entirely conserved between humans and these model organisms [3]. Finally, the quantity of ankyrin-like domain repeats varies very little between the organisms above [3].
Are there other NOTCH1 isoforms?
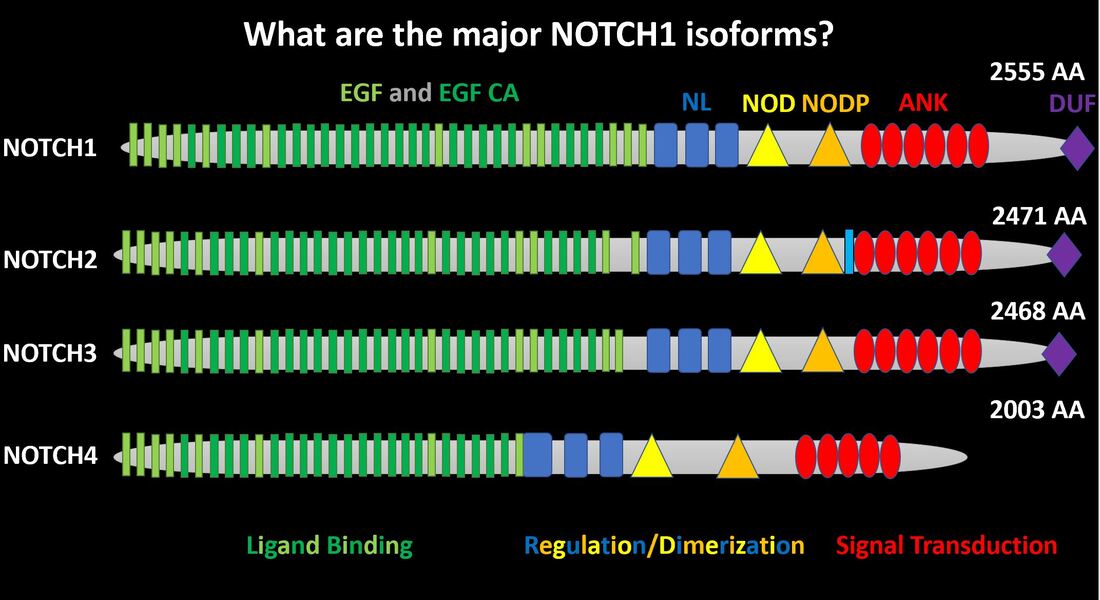
The NOTCH family contains four members that show considerable conservation between EGF, EGF CA, NL, NOD, NODP, and ANK domains [3]. NOTCH2 contains a small transmembrane domain (light blue) while NOTCH4 lacks a domain of unknown function (purple) [3].
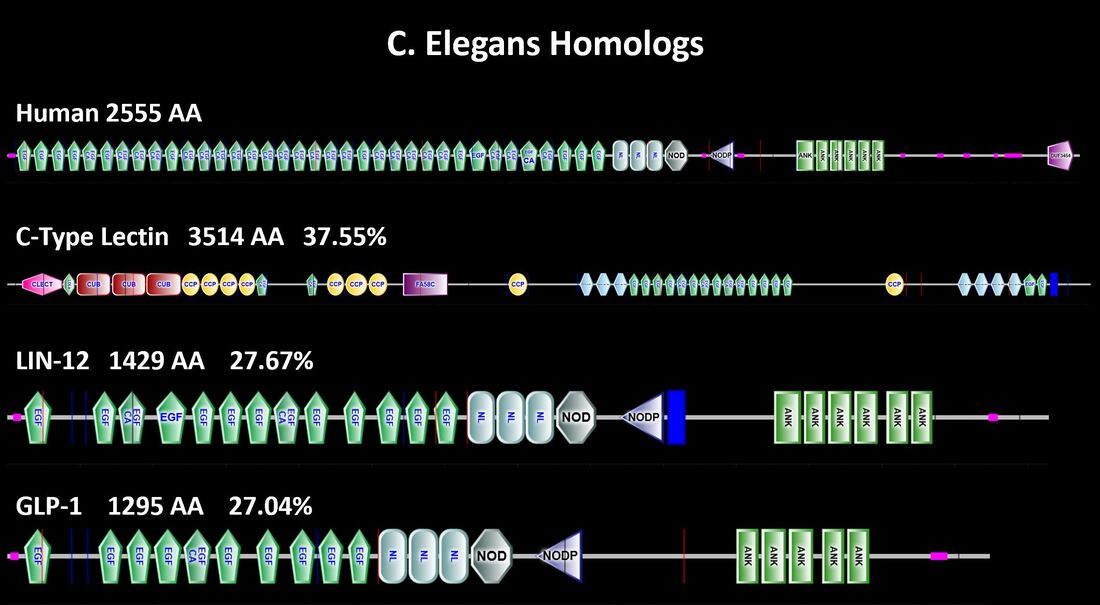
How does C. elegans compare?
As stated previously, C. elegans possesses three genes of considerable homology with NOTCH1 [3]. Although C-type lectin has the highest percent identicality it is considerably longer than NOTCH1 and contains several domains that are not in common [3].
Where are common NOTCH1 mutations?
I have identified four clinically relevant mutations thus far from primary literature [4]. Each occurs within the EGF and EGF calcium binding domain. I will update this as I progress through papers.
What are common NOTCH1 motifs?
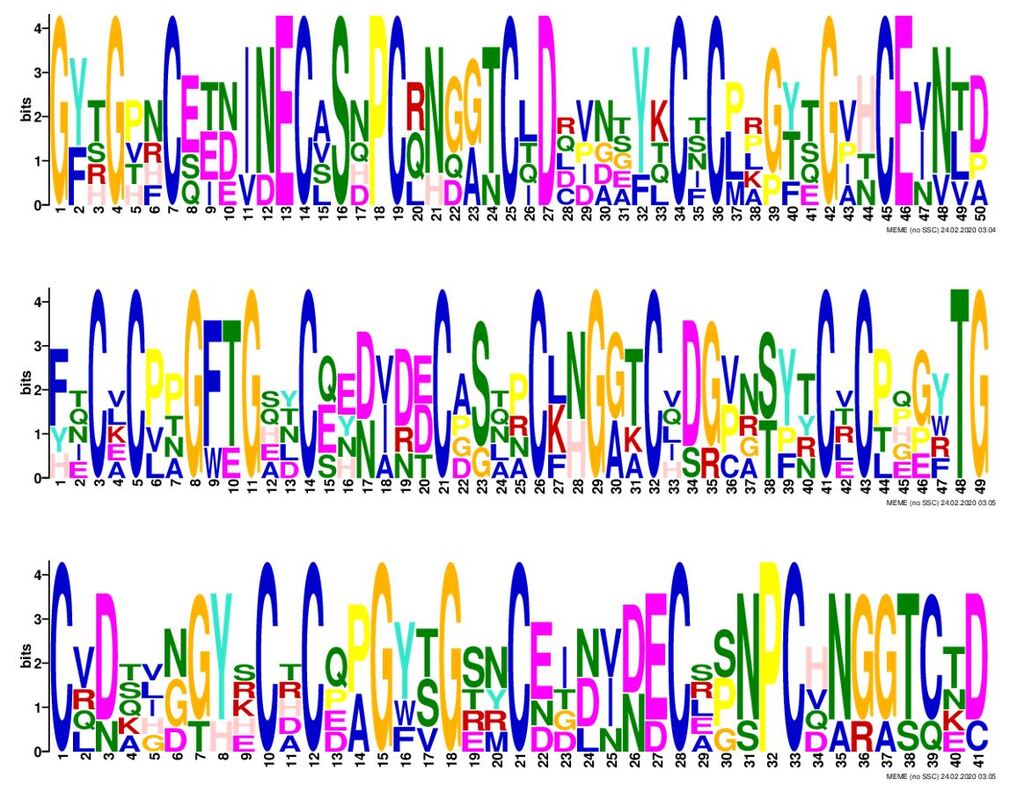
Above are the three most significant NOTCH1 motifs as determined by meme [5]. The considerable amount of cysteine residues within these motifs may indicate that disulfide bridging is important to NOTCH1 secondary, tertiary, or quaternary structure.
Conclusions
NOTCH has four isoforms that share the same domains for ligand binding, regulation/dimerization, and signal transduction. These domains are seen across each of the model organisms. Of importance is the EGF-like and EGF-like calcium binding domains of NOTCH1. These domains represent 36 repeats in humans and between 13-37 in the model organisms above. In addition, many of the common clinically defined mutations are found within this domain. As previously stated, a major phenotype of AoVD1 is aortic valve calcification. Do mutations in this region impair cellular calcium homeostasis and lead to the given phenotype? A final takeaway from domain analysis is the cysteine-rich motifs in NOTCH1. Where are these found? Do they contribute to protein folding? And does mutation of these residues play a role in AoVD1?