NOTCH1 Protein
What are protein interactions?
Biological processes depend not only on the proper amounts of protein produced in a cell but the proper interactions between these proteins as well. Characterization of an organism's "interactome" attempts to isolate which interactions occur because of functional biochemical properties while disregarding those that happen because of random chance [1].
How are protein interactions visualized?
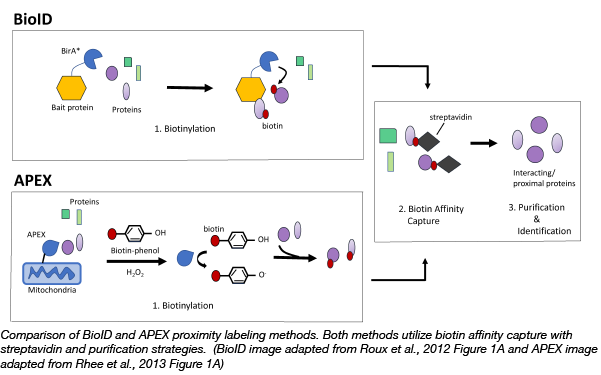
Two methods illustrating the characterization of protein-protein interactions are shown above. In BioID, a biotin ligase is incorporated into cells where it then biotinylates interacting proteins [2]. Through affinity capture, these biotinylated proteins can be isolated by streptavidin, purified, and finally determined by liquid chromatography/mass spectrometry [2]. This strategy is often utilized for interactions with insoluble or membrane-associated proteins [2]. APEX, on the other hand, fuses interacting proteins to a peroxidase [3]. Upon treatment with hydrogen peroxide and biotin-phenol these proteins are labeled with biotin as well [3]. Purification and characterization can then be carried out in much the same way as seen in BioID [3].
What does NOTCH1 interact with?
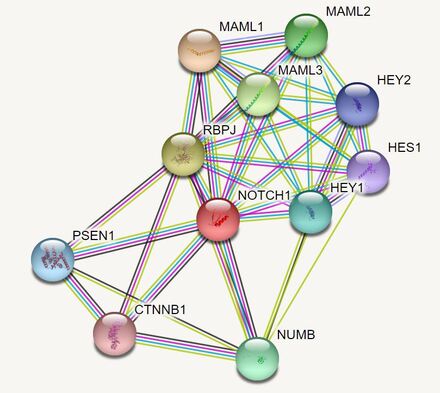
Human NOTCH1 interacts with a set of transcription factors and transcriptional co-activators (MAML1, MAML2, MAML3, HES1, HEY1, HEY2, RBPJ) which act most commonly in complex with histone acetylases and deacetylases to regulate enhancer elements of cellular fate [6-9]. NUMB encodes a membrane-bound protein key in neurodevelopment [10], CTNNB1 encodes catenin beta which makes up adheren junctions [11], and PSEN1 encodes presenilin which cleaves the NOTCH receptor [12].
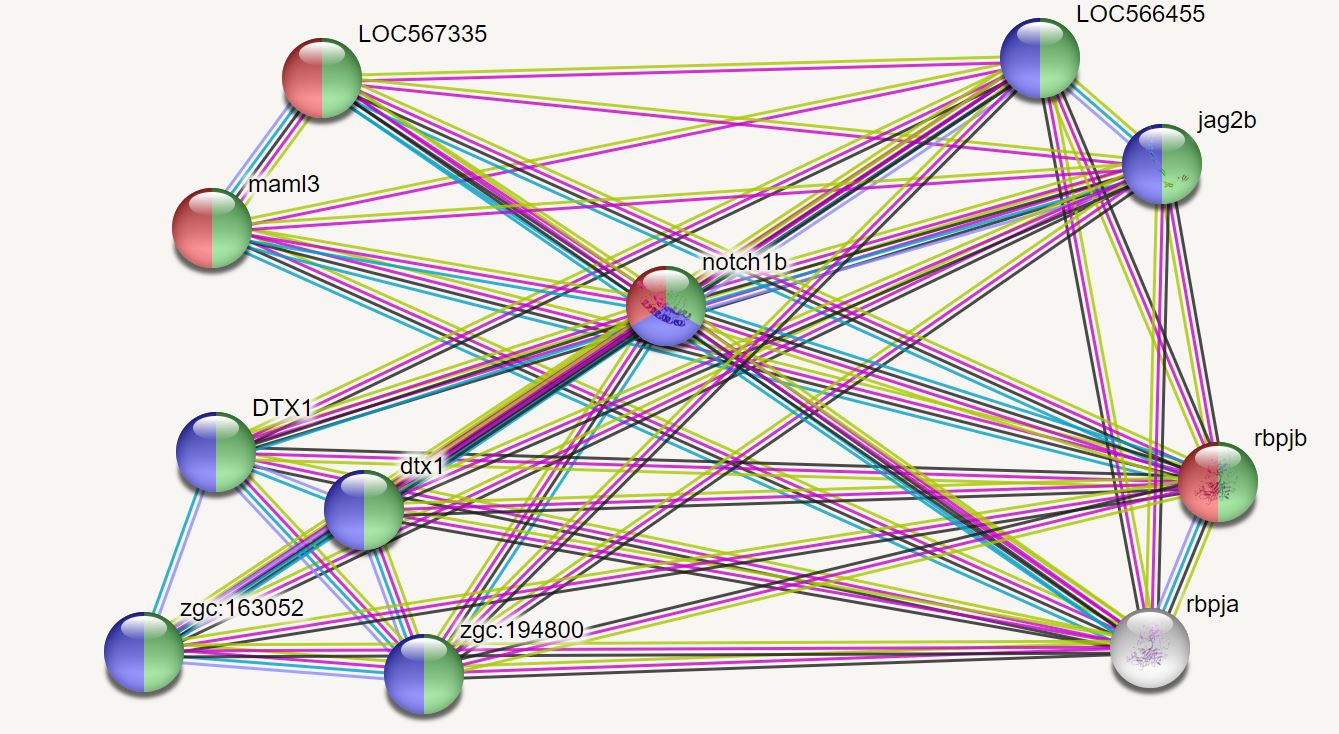
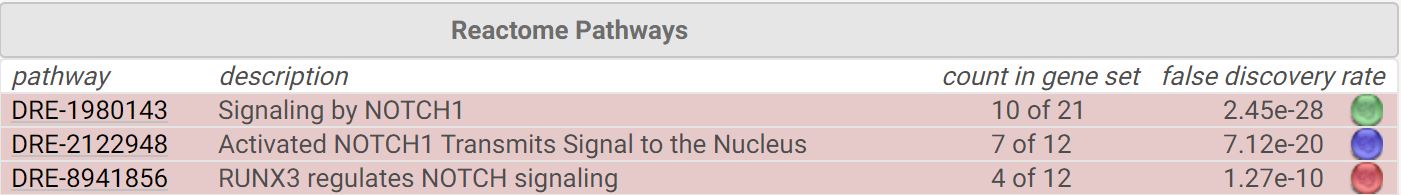
The zebrafish interaction network can be divided into two main functions, those that contribute to nuclear transmission and those that work together with RUNX3 [13], a DNA binding protein, to regulate NOTCH signaling.
|
JAG2B
|
RBPJA, RBPJB
|
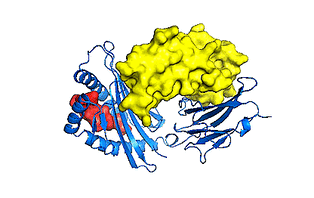
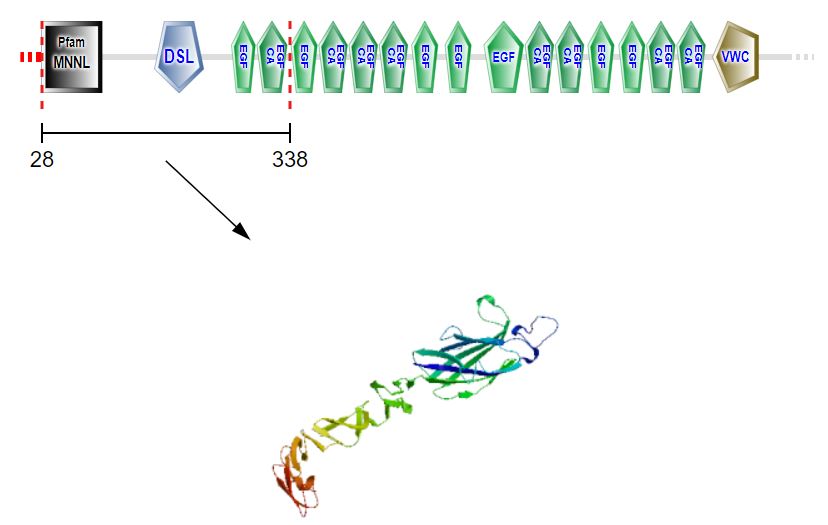
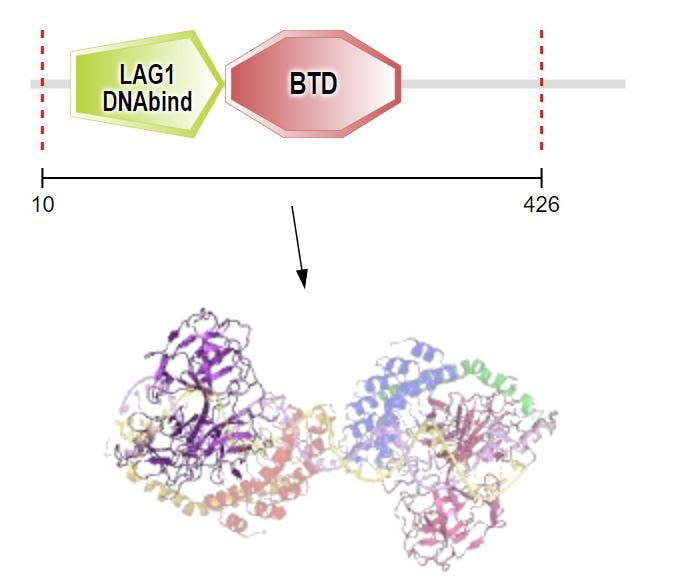
Pictured above are the domains/protein structures of the zebrafish interactome.
Conclusion
STRING interaction network analysis reveals that human NOTCH1 interacts within a complex transcriptional network that controls cell fate determination. In addition to this, it is cleaved by presenilin and interacts at the cell surface with NUMB and CTNNB1. While the zebrafish network is less defined, it can easily be categorized by the roles interactors play within the cellular environment. It also reveals that RUNX3 may be a key player in NOTCH signaling regulation.