NOTCH1 Gene
What is chemical genomics?
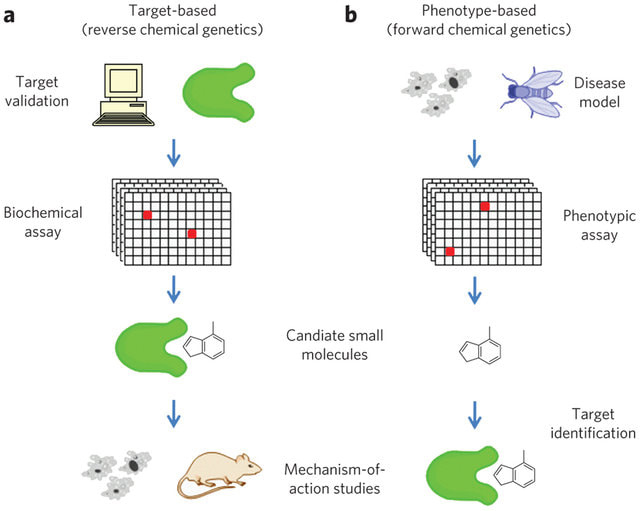
Chemical genomics takes advantage of small molecules and antimicrobial agents with known properties to identify future drug targets [1]. It can also be used on compounds of unknown property to understand toxicity [1]. Forward genetic approaches to chemical genomics take advantage of a known phenotype induced by drug application in the attempt of isolating the sequence change induced [1,2]. On the other hand, reverse genetics starts at the sequence of interest, introduces change upon drug administration, and looks to isolate the mutant phenotype induced by this change [1,2].
What inhibits NOTCH1?
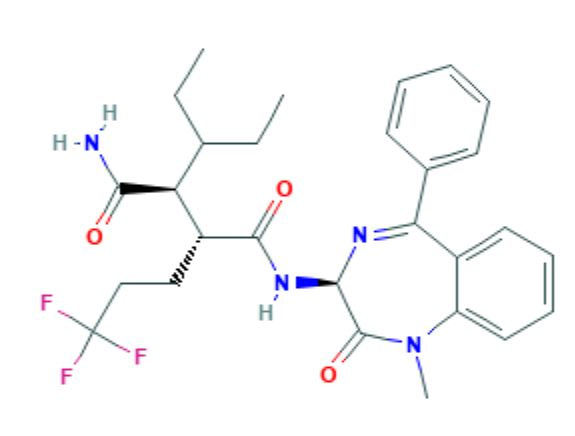
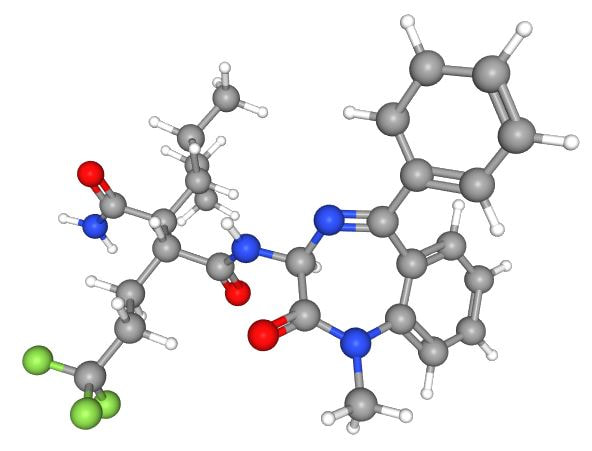
The 2D and 3D structure of (2S,3R)-N'-[(3S)-1-Methyl-2-oxo-5-phenyl-3H-1,4-benzodiazepin-3-yl]-2-pentan-3-yl-3-(3,3,3-trifluoropropyl) butanediamide can be found above [3]. This is one of the three most potent small molecule inhibitors of human NOTCH1 (Activity value : IC50 0.001 uM) [3]. It has a molecular weight of 530.6 g/mol and its chemical formula is C28H33F3N4O3 [3]. Two similar compounds with the chemical formulas C29H25F5N4O3 and C30H26F6N4O3 have the same activity value of IC50 0.001 uM [4,5].
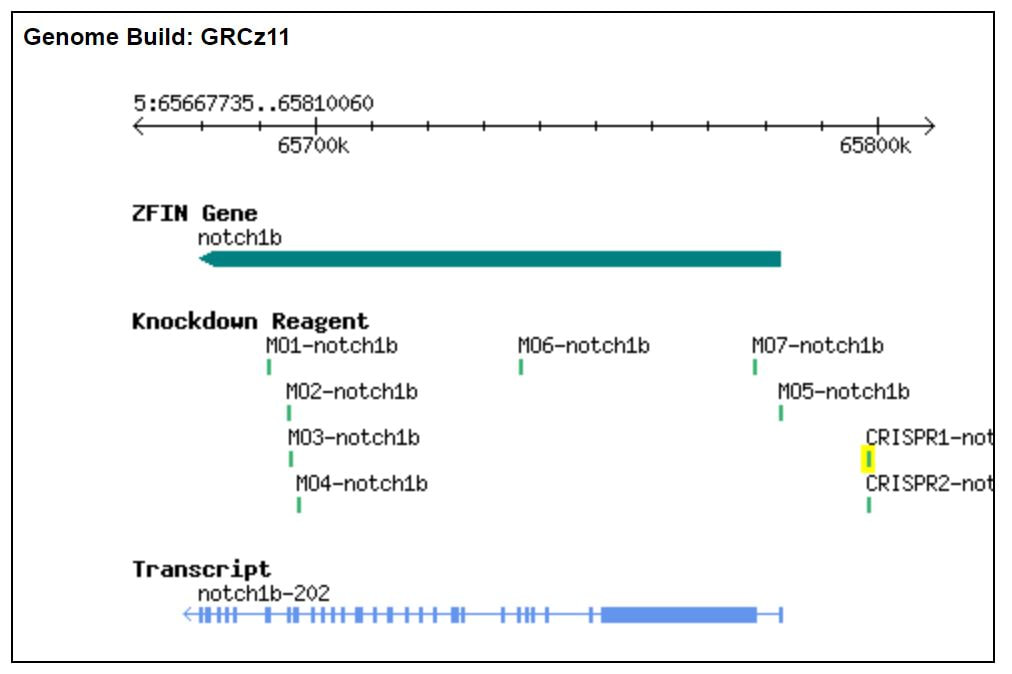
Reagents for, and analysis of, NOTCH1 modification can be found on ZFIN [6]. Above is a gene/transcript map with featured splice blocking morpholino and CRISPR reagents for the knockdown of NOTCH1b, the zebrafish homolog of NOTCH1 [6].
Conclusion
It has been documented that NOTCH1 can be inhibited by small molecules as well as more specific modifiers like splice blocking morpholinos and CRISPR. These tools present the possibility of NOTCH1 knockdown to study aberrant aortic valve formation.